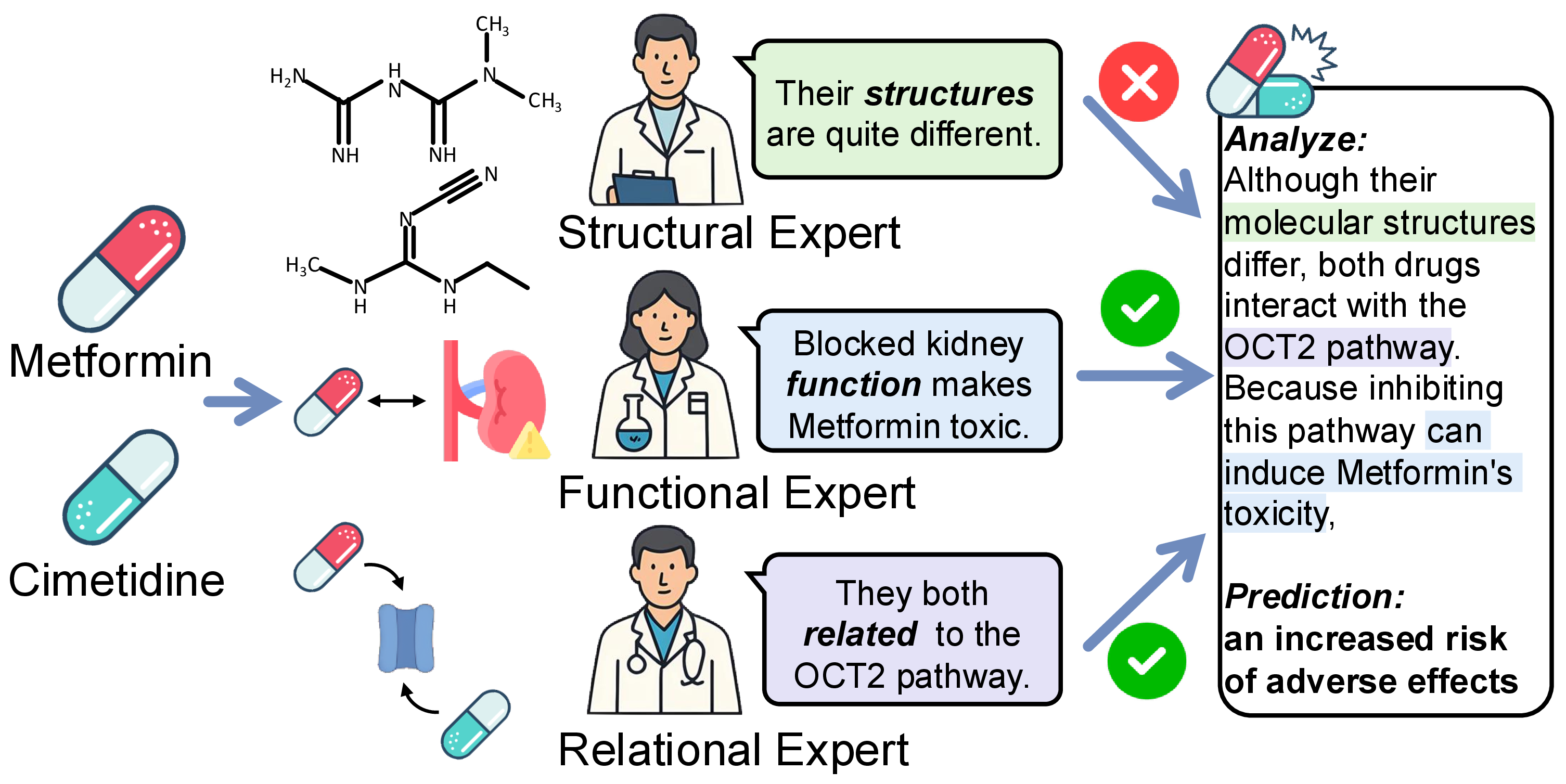
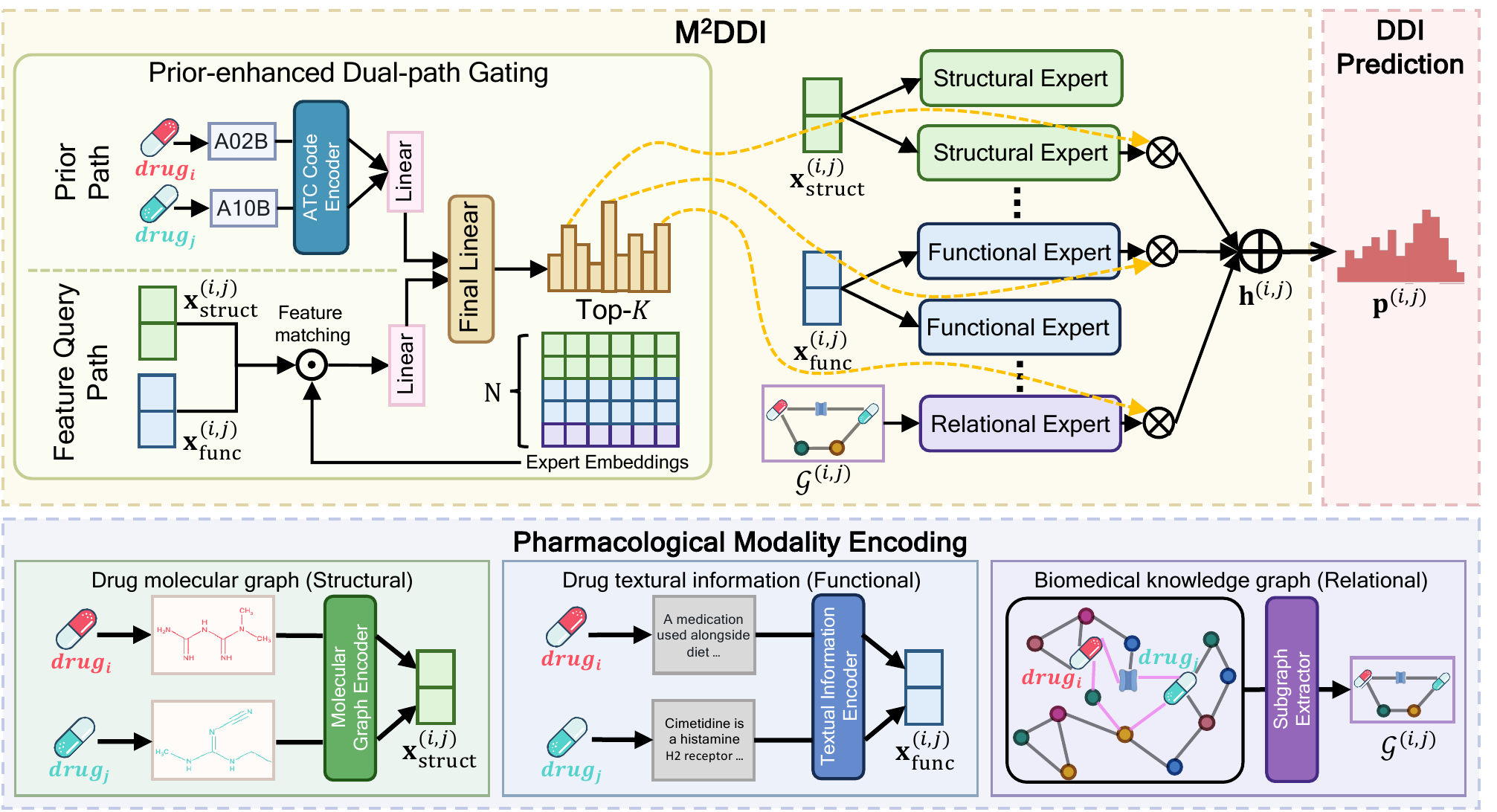
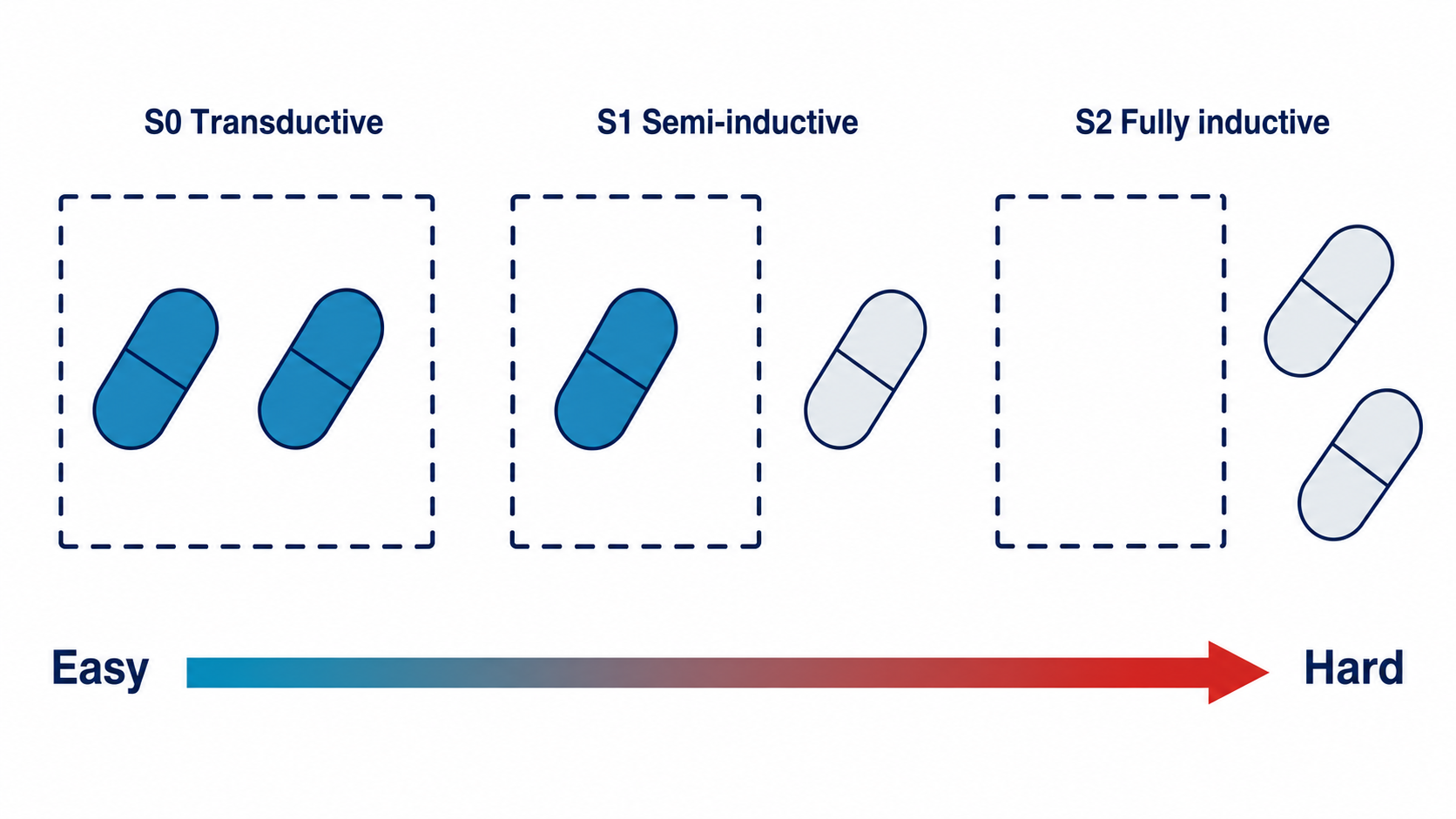
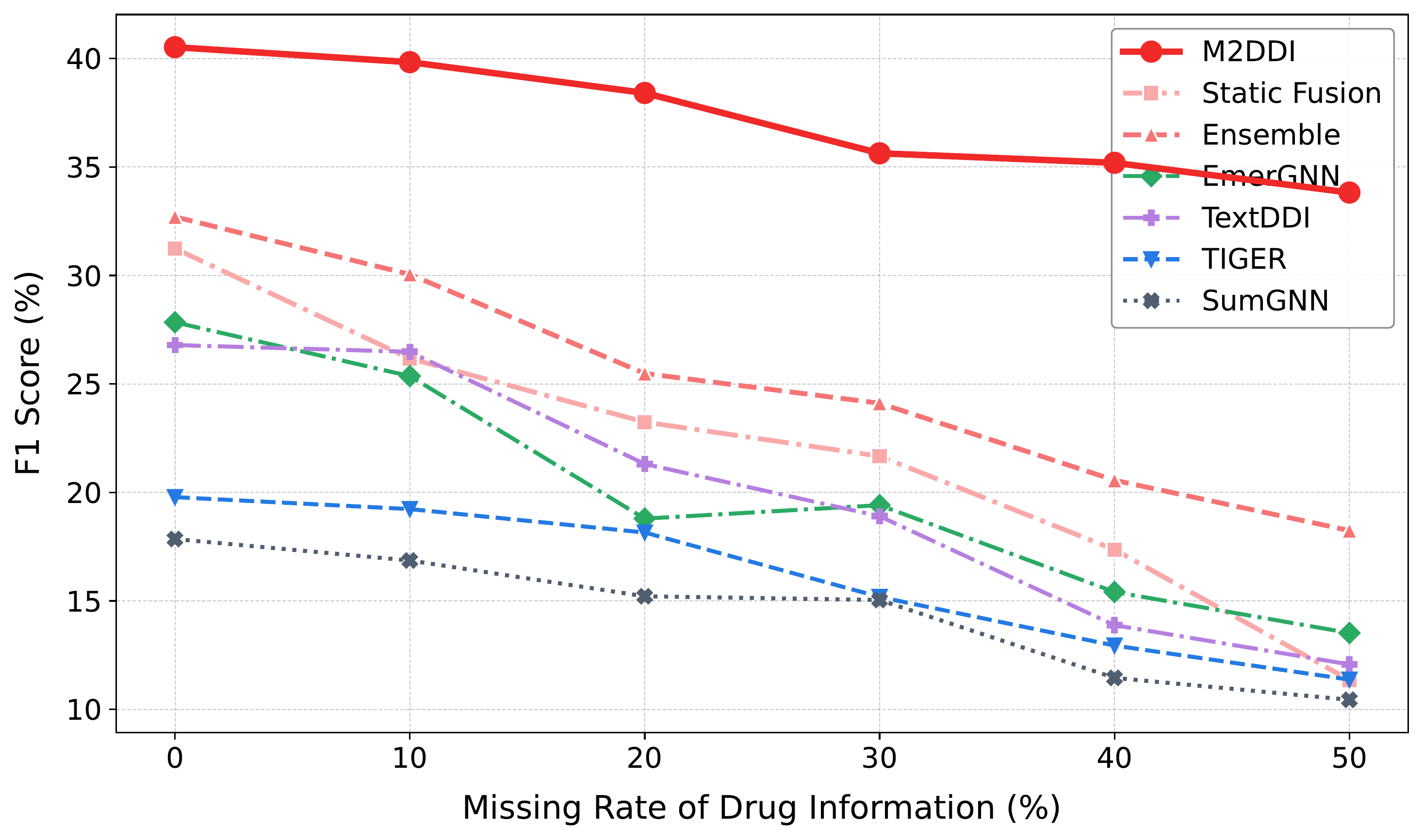
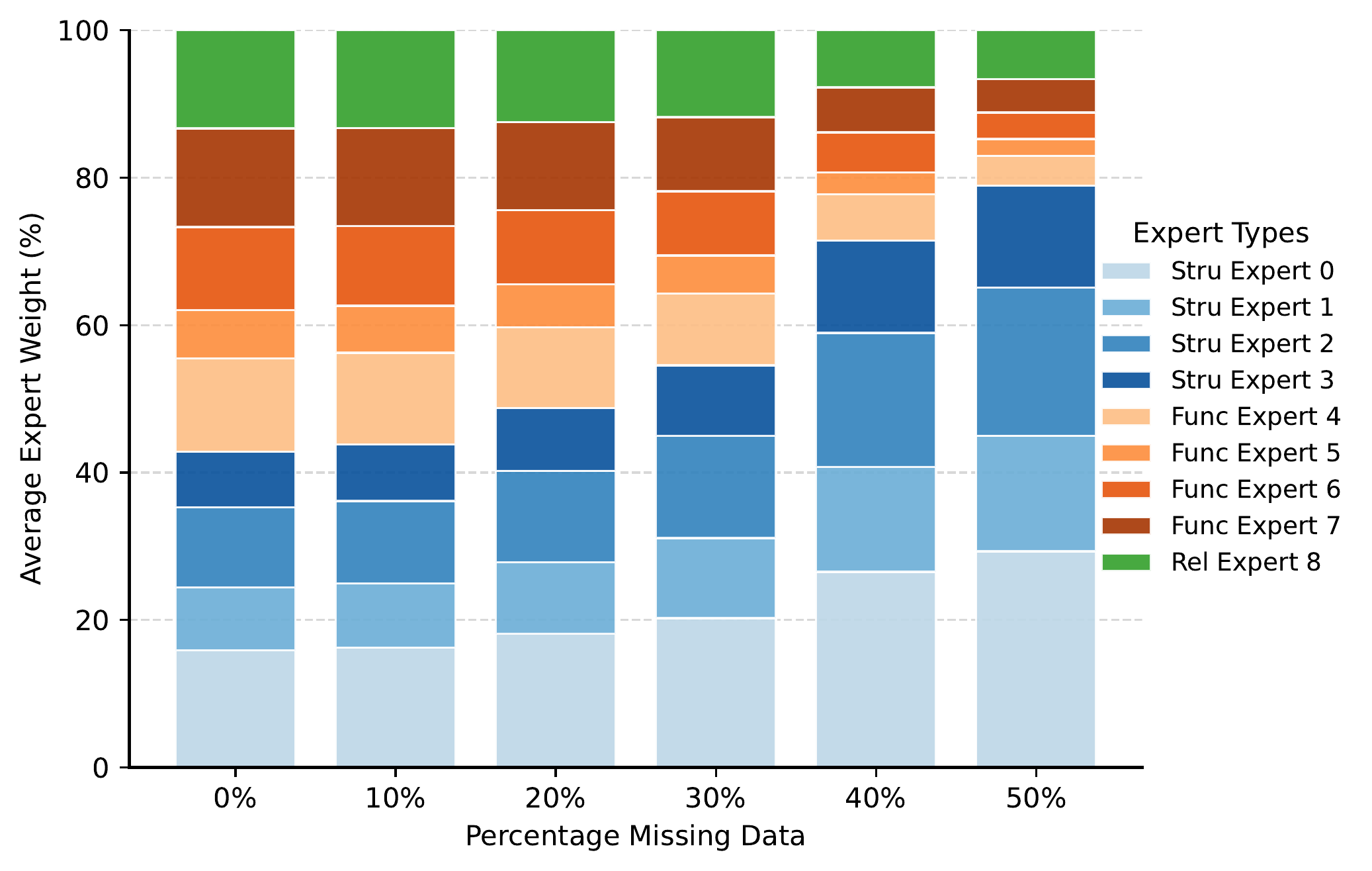
Molecular learning sits at the intersection of chemistry, biology, and machine learning.
The goal is to computationally understand molecular structure, predict molecular properties, and reason about molecular behavior.
Core Question
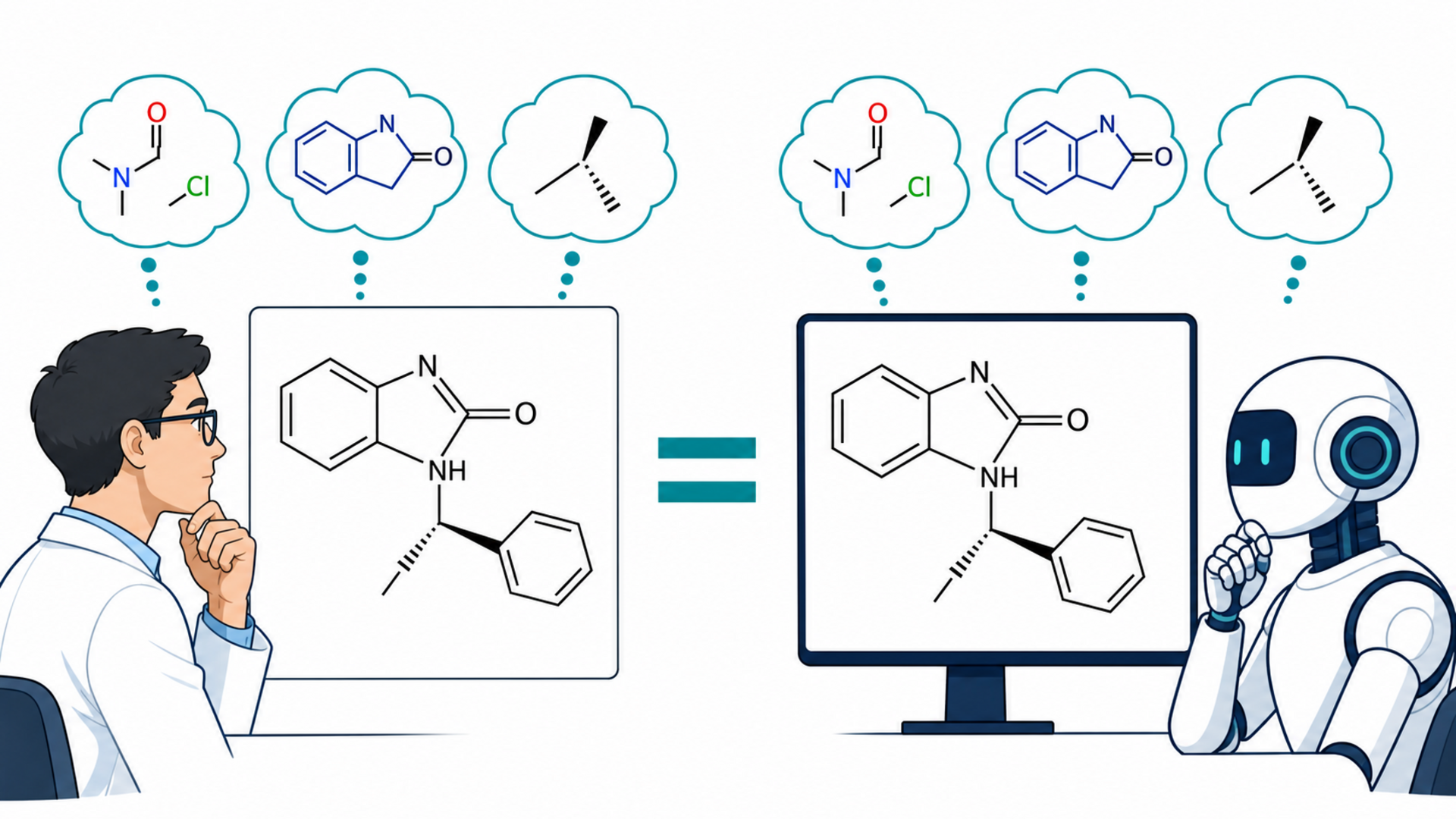
How can we build computational systems that understand molecules as well as expert chemists do?